Chapter 10: Introduction to Metabolism – Enzymes and Energy

Chapter Outline
- 10.1 Energy Changes During Chemical Reactions
- 10.2 Enzymes
- 10.3 Energy Transfer in Living Systems: Adenosine Triphosphate (ATP)
- 10.4 Energy Transfer in Living Systems: Electron Transfer
- 10.5 Metabolic Pathways
Introduction
In Part 4 of this text, we will address how living things manage their energy needs. Many autotrophs (organisms able to form biological macromolecules from simple inorganic compounds) are photosynthetic, and obtain energy from light. Both autotrophs and heterotrophs (organisms that have to eat complex organic substances to live) can break down biological macromolecules to release useable energy. Consider the oxidation of one molecule of glucose:
C6H12O6 + 6 O2 → 6 CO2 + 6 H2O
If this reaction occurs in a single step, combustion occurs, with energy released as the light and heat of a flame. Obviously, this single-step reaction is not compatible with life. Fortunately, evolution has resulted in a slower way to release energy from glucose and harness that energy in a safe form for the cell’s use: ATP.
Meeting a cell’s energy needs is one of the functions of metabolism. Technically speaking, metabolism is the sum of all reactions occurring in a living organism. More explicitly, metabolism deals with how an organism manages its material and energy resources, including:
-
Breaking down nutrients
-
Generating & storing energy
-
Synthesizing cellular building blocks (macromolecules)
-
Eliminating waste & potentially harmful substances
In this course, we focus on the first two roles of metabolism, and only a portion of those. If you take an advanced biochemistry course, you will delve further into the pathways that allow organisms to function. To understand the basics of metabolism, you need an understanding of how living things manage energy and how enzymes allow them to do so.
10.1 | Energy Changes During Chemical Reactions
Learning Objectives
By the end of this section, you will be able to:
- Identify endergonic and exergonic reactions and describe the change in Gibbs free energy.
- Define activation energy and explain why even exergonic reactions have an activation energy.
10.1.1 Gibbs Free Energy
It is critical for scientists to be able to quantify and express the energy changes associated with chemical reactions, specifically so that the energy released from one reaction be compared to that of another reaction. A measurement of Gibbs free energy is used to quantitate these energy transfers. Gibbs free energy (abbreviated with the letter G) was named after Josiah Willard Gibbs, the scientist who developed the measurement.
Recall that according to the second law of thermodynamics, all energy transfers involve the loss of some amount of energy in an unusable form such as heat, resulting in entropy. Gibbs free energy specifically refers to the energy associated with a chemical reaction that is available after entropy is accounted for. In other words, Gibbs free energy is usable energy, or energy that is available to do work.
Every chemical reaction involves a change in free energy, called delta G (∆G). The change in free energy can be calculated for any system that undergoes a change, such as a chemical reaction. To calculate ∆G, start with the total energy change of the system, which is called enthalpy, and is denoted as ΔH. Then subtract the amount of energy that is lost to entropy (denoted as ∆S). The formula for calculating ∆G is as follows, where the symbol T refers to absolute temperature in Kelvin (degrees Celsius + 273):
ΔG = ΔH − TΔS
The standard free energy change of a chemical reaction is expressed either in kilojoules per mole (kJ/mol) or kilocalories per mole (kcal/mol) under standard pH, temperature, and pressure conditions. In biological systems, standard conditions are typically pH 7.0, 25 degrees Celsius, and 1 atm pressure. However, cellular conditions can vary considerably from these standard conditions, and so actual ∆G values for biological reactions may differ.
10.1.2 Endergonic Reactions and Exergonic Reactions
If energy is released during a chemical reaction, then the resulting value from the above equation for ΔG will be a negative number. A negative ∆G means that the products of the reaction have less free energy than the reactants, because they gave off some free energy during the reaction. Reactions that have a negative ∆G and consequently release free energy are called exergonic reactions. (Think: exergonic means energy is exiting the system.) These reactions are also referred to as spontaneous reactions, because they can occur without the addition of energy into the system (Figure 10.2). Understanding which chemical reactions are spontaneous and release free energy is extremely useful for biologists, because these reactions can be harnessed to perform work inside the cell. An important distinction must be drawn between the term spontaneous and the idea of a chemical reaction that occurs immediately. Contrary to the everyday use of the term, a spontaneous reaction is not one that suddenly or quickly occurs. The rusting of iron is an example of a spontaneous reaction that occurs slowly, little by little, over time.
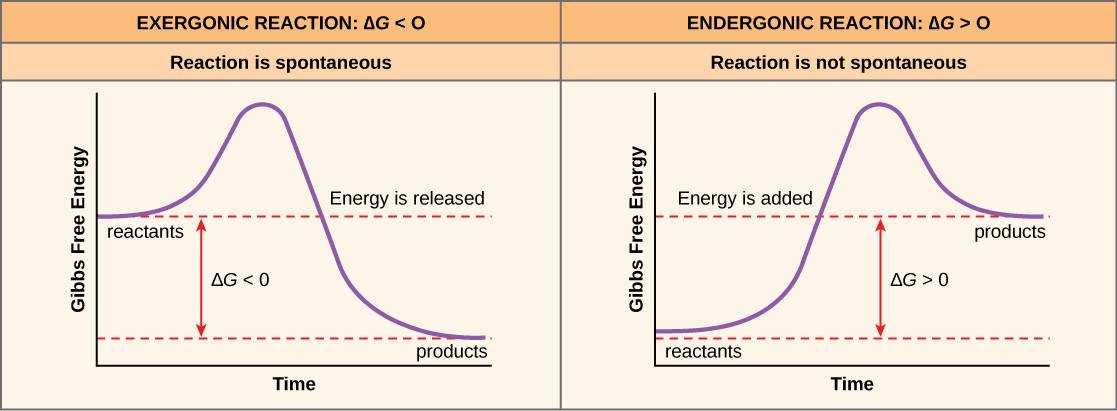
If a chemical reaction requires an input of energy rather than releasing energy, then the ∆G for that reaction will be a positive value. In this case, the products have more free energy than the reactants. Thus, the products of these reactions can be thought of as energy-storing molecules. These chemical reactions are called endergonic reactions, and they are non-spontaneous. An endergonic reaction will not take place on its own with the addition of energy (Figure 10.2).
Let’s consider the example of the synthesis and breakdown of the food molecule, glucose. In metabolism, the building of complex molecules, such as sugars, from simpler ones is called an anabolic process and requires energy. Therefore, the chemical reactions involved in anabolic processes are endergonic reactions. On the other hand, the catabolic process of breaking sugar down into simpler molecules releases energy in a series of exergonic reactions. Figure 10.3 shows some other examples of endergonic and exergonic reactions.

Another important concept in the study of metabolism and energy is that of chemical equilibrium. Most chemical reactions are reversible. They can proceed in both directions, releasing energy into their environment in one direction, and absorbing it from the environment in the other direction. The same is true for the chemical reactions involved in cell metabolism, such as the breaking down and building up of proteins into and from individual amino acids, respectively. Reactants within a closed system will undergo chemical reactions in both directions until a state of equilibrium is reached. This state of equilibrium is one of the lowest possible free energy and a state of maximal entropy.
10.1.3 Activation Energy
Even exergonic reactions require a small amount of energy input to get going before they can proceed with their energy-releasing steps. These reactions have a net release of energy, but still require some energy in the beginning. This small amount of energy input necessary for all chemical reactions to occur is called the activation energy.
Why would an energy-releasing, negative ∆G reaction actually require some energy to proceed? The reason lies in the steps that take place during a chemical reaction. During chemical reactions, certain chemical bonds are broken and new ones are formed. For example, when a glucose molecule is broken down, bonds between the carbon atoms of the molecule are broken. Since these are energy-storing bonds, they release energy when broken. However, to get them into a state that allows the bonds to break, the molecule must be somewhat contorted. A small energy input is required to achieve this contorted state. This contorted state is called the transition state, and it is a high-energy, unstable state. For this reason, reactant molecules don’t last long in their transition state, but very quickly proceed to the next steps of the chemical reaction.
Free energy diagrams (also called reaction coordinate diagrams) illustrate the energy profiles for a given reaction (Figure 10.4). Whether the reaction is exergonic or endergonic determines whether the products of the reaction will exist at a lower or higher energy state than the reactants. However, regardless of this measure, the transition state will always exists at a higher energy state than the reactants.
Where does the activation energy required by chemical reactants come from? The source of the activation energy needed to push reactions forward is typically heat energy from the surroundings. Heat energy speeds up the motion of molecules, increasing the frequency and force with which they collide; it also moves atoms and bonds within the molecule slightly, helping them reach their transition state. For this reason, heating up a system will cause chemical reactants within that system to react more frequently. Once reactants have absorbed enough heat energy from their surroundings to reach the transition state, the reaction will proceed.
The activation energy of a particular reaction determines the rate at which it will proceed. The higher the activation energy, the slower the chemical reaction will be. The example of iron rusting illustrates an inherently slow reaction. This reaction occurs slowly over time because of its high activation energy. Additionally, the burning of many fuels, which is strongly exergonic, will take place at a negligible rate unless their activation energy is overcome by sufficient heat from a spark. Once they begin to burn, however, the chemical reactions release enough heat to continue the burning process, supplying the activation energy for surrounding fuel molecules.
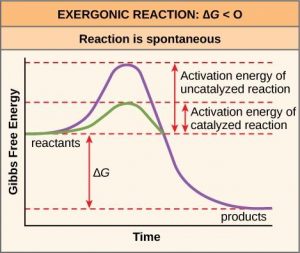
Like these reactions outside of cells, the activation energy for most cellular reactions is too high for heat energy to overcome at efficient rates. In other words, in order for important cellular reactions to occur at appreciable rates (number of reactions per unit time), their activation energies must be lowered (Figure 10.4). As we are about to discuss, enzymes function by lowering the activation energy and increasing the reaction rate of reactions. This is a very good thing as far as living cells are concerned. Important macromolecules, such as proteins, DNA, and RNA, store considerable energy, and their breakdown is exergonic. If cellular temperatures alone provided enough heat energy for these exergonic reactions to overcome their activation barriers, the essential components of a cell would disintegrate.
10.2 | Enzymes
Learning Objectives
By the end of this section, you will be able to:
- Explain how enzymes function as molecular catalysts.
- Discuss enzyme regulation by various factors.
A substance that speeds up a chemical reaction without being changed by that reaction is a catalyst. Proteins that catalyze biochemical reactions are called enzymes. An example of an enzyme is salivary amylase, which hydrolyzes its substrate amylose, a component of starch.
Enzymes perform the critical task of lowering the activation energies of chemical reactions inside the cell. Enzymes do this by binding to the reactant molecules, and holding them in such a way as to make the chemical bond-breaking and bond-forming processes take place more readily. Since all enzymes increase the rate of reaction, they are considered to be organic catalysts (Figure 10.4). Note that although enzymes can speed up reactions, they can never change an endergonic reaction to an exergonic one. Whether a reaction is endergonic or exergonic depends only on the chemical makeup on the reactant and product molecules and cannot be changed by a catalyst.
10.2.1 Enzyme Active Site and Substrate Specificity
The chemical reactants to which an enzyme binds are called the enzyme’s substrates. There may be one or more substrates, depending on the particular chemical reaction. In some reactions, a single-reactant substrate is broken down into multiple products. In others, two substrates may come together to create one larger molecule.
The location within the enzyme where the substrate binds is called the enzyme’s active site. Since enzymes are proteins, there is a unique combination of amino acid side chains, which creates a very specific chemical environment within the active site. This specific environment is suited to bind to a specific substrate (or substrates). Due to this jigsaw puzzle-like match between an enzyme and its substrates, enzymes are known for their specificity. The “best fit” results from the shape and the amino acid functional group’s attraction to the substrate. There is a specifically matched enzyme for each substrate and, thus, for each chemical reaction.
Since enzymes are proteins, their shape is sensitive to variations in temperature and pH. Increasing or decreasing the temperature outside of an optimal range can affect chemical bonds within the active site in such a way that they are less well suited to bind substrates. High temperatures will eventually cause enzymes to denature. Likewise, pH can also affect enzyme function. Enzymes are suited to function best within a certain pH range, and extreme pH values can cause enzymes to denature.
Induced Fit and Enzyme Function
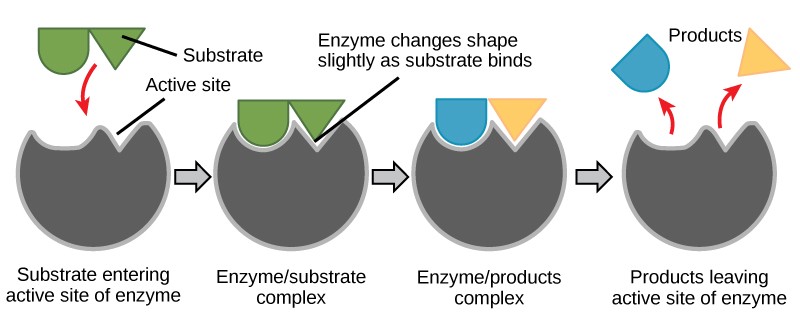
For many years, scientists thought that enzyme-substrate binding took place in a simple “lock-and-key” fashion. This model asserted that the enzyme and substrate fit together perfectly in one instantaneous step. However, current research supports a more refined view called induced fit (Figure 10.5). The induced-fit model expands upon the lock-and-key model by describing a more dynamic interaction between enzyme and substrate. As the enzyme and substrate come together, their interaction causes a mild shift in the enzyme’s structure that confirms an ideal binding arrangement between the enzyme and the transition state of the substrate. This ideal binding maximizes the enzyme’s ability to catalyze its reaction.
Enzymes will always return to their original state once the reaction is complete. One of the hallmark properties of enzymes is that they remain ultimately unchanged by the reactions they catalyze. After an enzyme is finished catalyzing a reaction, it releases its product(s).
10.2.2 Control of Metabolism Through Enzyme Regulation
It would seem ideal to have a scenario in which all of the enzymes encoded in an organism’s genome existed in abundant supply and functioned optimally under all cellular conditions, in all cells, at all times. In reality, this is far from the case. A variety of mechanisms ensure that this does not happen. Cellular needs and conditions vary from cell to cell, and change within individual cells over time. The required enzymes and energetic demands of stomach cells are different from those of fat storage cells, skin cells, blood cells, and nerve cells. Furthermore, a digestive cell works much harder to process and break down nutrients during the time that closely follows a meal compared with many hours after a meal. As these cellular demands and conditions vary, so do the amounts and functionality of different enzymes.
The relative amounts and functioning of the variety of enzymes within a cell ultimately determine which reactions will proceed and at which rates. This determination is tightly controlled. In certain cellular environments, enzyme activity is partly controlled by environmental factors, like pH and temperature. There are other mechanisms through which cells control the activity of enzymes and determine the rates at which various biochemical reactions will occur.
Regulation of Enzymes by Small Molecules
There are many different kinds of molecules that inhibit or promote enzyme function. In some cases, an inhibitor molecule binds to the active site of an enzyme and physically blocks the substrate from binding. This is called competitive inhibition because an inhibitor molecule competes with the substrate for active site binding (Figure 10.6).
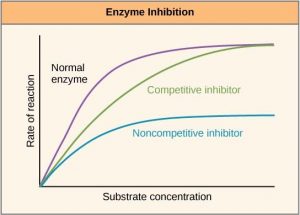
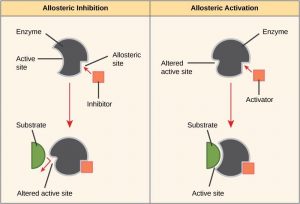
In other cases, an inhibitor molecule binds to the enzyme in a location other than the active site, called the allosteric site, and causes a shape change in the enzyme that blocks substrate binding to the active site. This type of inhibition is called allosteric inhibition (Figure 10.7). There are also allosteric activators, which bind to an allosteric site and induce a conformational change that increases the affinity of the enzyme for its substrate(s).

Drug Discovery by Looking for Inhibitors of Key Enzymes in Specific Pathways
Enzymes are key components of metabolic pathways. Understanding how enzymes work and how they can be regulated is a key principle behind the development of many of the pharmaceutical drugs (Figure 10.8) on the market today. Biologists working in this field collaborate with other scientists, usually chemists, to design drugs.
Consider statins for example—which is the name given to the class of drugs that reduces cholesterol levels. These compounds are essentially inhibitors of the enzyme HMG-CoA reductase. HMG-CoA reductase is the enzyme that synthesizes cholesterol from lipids in the body. By inhibiting this enzyme, the levels of cholesterol synthesized in the body can be reduced. Similarly, acetaminophen, popularly marketed under the brand name Tylenol, is an inhibitor of the enzyme cyclooxygenase. While it is effective in providing relief from fever and inflammation (pain), its mechanism of action is still not completely understood.
How are drugs developed? One of the first challenges in drug development is identifying the specific molecule that the drug is intended to target. In the case of statins, HMG-CoA reductase is the drug target. Drug targets are identified through painstaking research in the laboratory. Identifying the target alone is not sufficient; scientists also need to know how the target acts inside the cell and which reactions go awry in the case of disease. Once the target and the pathway are identified, then the actual process of drug design begins. During this stage, chemists and biologists work together to design and synthesize molecules that can either block or activate a particular reaction. However, this is only the beginning: both if and when a drug prototype is successful in performing its function, then it must undergo many tests from in vitro experiments to clinical trials before it can get FDA approval to be on the market.
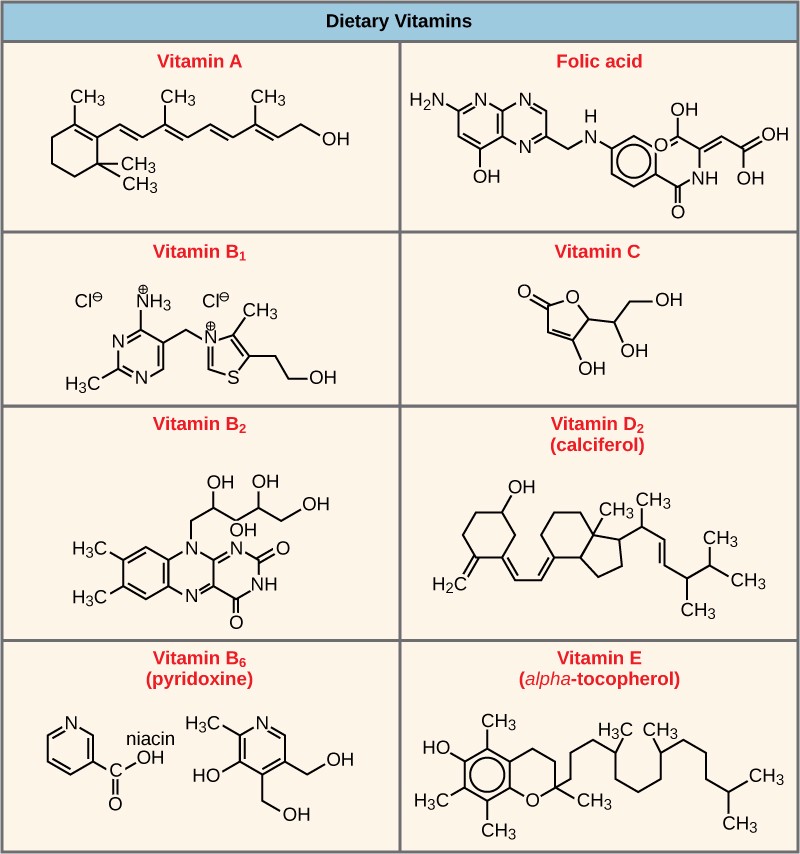
Many enzymes don’t work optimally, or even at all, unless bound to other specific non-protein helper molecules. Two types of helper molecules are cofactors and coenzymes. When these molecules bind to their enzymes, they promote optimal conformation and function. Cofactors are inorganic ions such as iron (Fe++) and magnesium (Mg++). Coenzymes are organic helper molecules. The most common sources of coenzymes are dietary vitamins (Figure 10.9). For example, vitamin C is a coenzyme for multiple enzymes that take part in building the important connective tissue component, collagen. Therefore, enzyme function is, in part, regulated by an abundance of various cofactors and coenzymes, which are supplied primarily by the diets of most organisms.
Regulation of Enzymes by Post-translational Modification
Some amino acid side chains in enzymes can be chemically modified after the protein itself is synthesized (translated from an mRNA). These changes are collectively called post-translational modifications (PTMs) (Table 10.1).
Table 10.1 Some common post-translational modifications.
| Name of modification | Chemical group added | Structure | Amino acids modified |
| Phosphorylation | Phosphate | PO34- | Serine, threonine, tyrosine |
| Glycosylation | Mono-, di-, and polysaccharides | Various | Asparagine (N-linked), serine or threonine (O-linked) |
| Ubiquitination | Ubiquitin | 8-kDa polypeptide of 76 amino acids | Lysine |
| Nitrosylation | Nitric oxide | NO | Cysteine |
| Methylation | Methyl | -CH3 | Many containing either O or N |
| Acetylation | Acetyl | -COCH3 | Lysine |
| Lipidation | Lipid | Many, but most well-known are prenyl and fatty acyl groups and glycosylphosphatidylinositols (GPI) | Various, including N- and C- termini |
Phosphorylation is the most well-studied PTM, where a phosphate group is added to a serine, threonine, or tyrosine residue by an enzyme known as a kinase. That phosphate group can then be removed by another enzyme known as a phosphatase.
How a post-translational modification affects a protein’s activity depends on a) the protein being modified, b) the PTM being added, and c) the site at which the PTM is added. Enzyme X might be activated by phosphorylation at serine 101, but deactivated by phosphorylation at serine 152. In this scenario, two different kinases would catalyze the addition of the phosphate group to the two different serine residues, and the kinases would likely respond to different cellular states.
Regulation of Enzymes by Changes in Gene Expression
As we will discuss in detail later in this text, all cells carry the same DNA blueprint, but individual cells express only a subset of those genes at a time. For instance, a cell growing in a low-serine environment might synthesize more of the enzyme molecules involved in the serine synthesis pathway than a cell growing in a serine-rich environment.
Regulation of Enzymes by Compartmentalization
In eukaryotic cells, molecules such as enzymes are usually compartmentalized into different organelles. This allows for yet another level of regulation of enzyme activity. Enzymes required only for certain cellular processes can be housed separately along with their substrates, allowing for more efficient chemical reactions. Examples of this sort of enzyme regulation based on location and proximity include the enzymes involved in the latter stages of cellular respiration, which take place exclusively in the mitochondria, and the enzymes involved in the digestion of cellular debris and foreign materials, located within lysosomes.
Table 10.2 Summary of enzyme regulation methods
| Method | Speed | Reversible? |
| Small molecules | Fast | Yes |
| Post-translational modifications | Fast | Yes |
| Gene expression | Slow | Yes, but on a longer timescale |
| Compartmentalization | N/A | No |
10.3 | Energy Transfer in Living Systems: Adenosine Triphosphate (ATP)
Learning Objectives
By the end of this section, you will be able to:
- Explain the role of ATP as the cellular energy currency.
- Describe how energy is released through hydrolysis of ATP.
10.3.1 ATP Structure & Function
As discussed above, even exergonic reactions require a small amount of activation energy in order to proceed. However, consider endergonic reactions, which require much more energy input, because their products have more free energy than their reactants. Within the cell, where does energy to power such reactions come from? The answer frequently lies with an energy-supplying molecule called adenosine triphosphate, or ATP. ATP is a small, relatively simple molecule (Figure 10.10), but, due to its chemical structure, it contains the potential for a quick burst of energy that can be harnessed to perform cellular work. ATP is the primary energy currency of cells and is used to power the majority of energy-requiring cellular reactions.
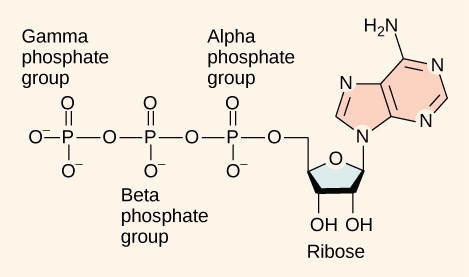
Adenosine triphosphate is comprised of a ribonucleotide containing adenine and three phosphate groups (Figure 10.10). When the bond that links the beta and gamma phosphate groups is broken, the reaction releases sufficient energy to power a variety of cellular reactions and processes.
The reason that this reaction releases so much energy is because the products of such bond breaking—adenosine diphosphate (ADP) and one inorganic phosphate group (Pi)—have considerably lower free energy than the reactants: ATP and a water molecule. Because this reaction takes place with the use of a water molecule, it is considered a hydrolysis reaction. In other words, ATP is hydrolyzed into ADP in the following reaction:
ATP + H2 O → ADP + Pi + free energy
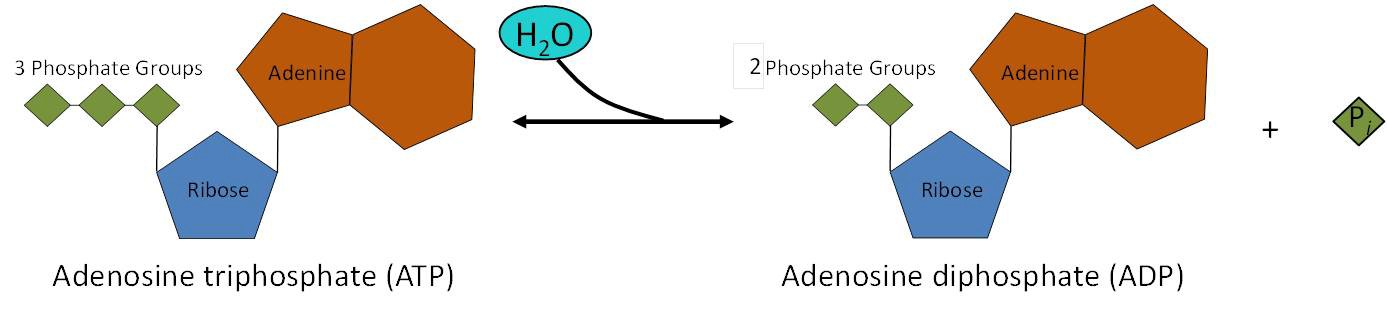
The same reaction is shown in Figure 10.11. Like most chemical reactions, the hydrolysis of ATP to ADP is reversible. The reverse reaction regenerates ATP + H2O from ADP + Pi.
ATP is a highly unstable molecule. Unless quickly used to perform work, ATP spontaneously dissociates into ADP + Pi, and the free energy released during this process is lost as heat. Cells use a process called energy coupling to harness the energy released by ATP hydrolysis to perform work inside the cell. In this process, an exergonic reaction, such as hydrolyzing ATP, is used to drive an endergonic reaction.
ATP is constantly being used and regenerated in a process known as the ATP cycle (Figure 10.12). ADP phosphorylation, an energy-requiring process, is coupled to a sufficiently energy-releasing process, nutrient breakdown, to allow ATP regeneration to occur. Then, ATP hydrolysis is coupled to cellular endergonic processes such as anabolism, active transport, and cell division, providing the required energy.

10.3.2 ATP Generation
Substrate-Level Phosphorylation
During the breakdown of glucose, a few ATP molecules are regenerated from ADP as a direct result of the chemical reactions that occur in the catabolic pathways. A phosphate group is removed from an intermediate reactant in the pathway, and the free energy of the reaction is used to add the third phosphate to an available ADP molecule, producing ATP (Figure 10.13). This very direct method of phosphorylation is called substrate-level phosphorylation.
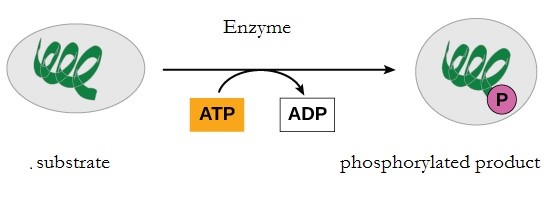
Oxidative Phosphorylation
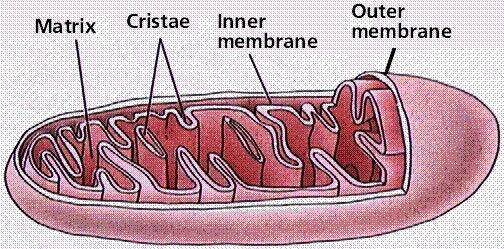
However, most of the ATP generated during glucose catabolism is derived from a much more complex process, chemiosmosis, which takes place in the mitochondria (Figure 10.14) of eukaryotic cells and in the plasma membrane of prokaryotic cells. The production of ATP by chemiosmosis in the mitochondria is called oxidative phosphorylation because it requires oxygen.
Chemiosmosis is also used in the light reactions of photosynthesis (in chloroplasts) to harness the energy of sunlight. This process is called photophosphorylation because of the involvement of light energy.
10.4 | Energy Transfer in Living Systems: Electron Transfer
Learning Objectives
By the end of this section, you will be able to:
- Discuss the importance of electrons in the transfer of energy in living systems.
- Identify reduction and oxidation reactions.
- Be familiar with common electron carriers.
10.4.1 The Carbon Cycle
The metabolism of sugar (a simple carbohydrate) is a classic example of the many cellular processes that use and produce energy. Living things consume sugar as a major energy source, because sugar molecules have a great deal of energy stored within their bonds. The breakdown of glucose into carbon dioxide and water is described by the equation:
C6H12O6 + 6O2 → 6CO2 + 6H2O + energy
Carbohydrates have their origins in photosynthesizing organisms, such as plants. During photosynthesis, plants use the energy of sunlight to convert carbon dioxide gas (CO2) into sugar molecules, such as glucose (C6H12O6). The synthesis of glucose is described by the reverse of the previous equation:
6CO2 + 6H2O + energy → C6H12O6 + 6O2
Because photosynthesis involves synthesizing a larger, energy-storing molecule (glucose), it requires an input of energy to proceed. First, energy from sunlight is transformed into chemical energy in the form of ATP. The stored energy in ATP is then used to build one molecule of glucose from six molecules of CO2.
Both ATP and glucose store energy in their chemical bonds. However, ATP is very unstable and cannot be stored for more than a few seconds. Therefore, photosynthetic cells use the ATP to make glucose, which is more stable and can be used to make disaccharides or polysaccharides for long-term energy storage.
When cells need to use this energy to do work, sugar molecules are broken down through a complex series of chemical reactions in order to harvest the energy stored in their chemical bonds. The energy harvested is used to make ATP molecules, which can be used to power many chemical reactions and processes in the cell. If ATP is the dollar bill of the cell’s energy currency, carbohydrates synthesized from glucose can be thought of as one of the organism’s savings accounts.
Together, photosynthesis and cellular respiration make up the carbon cycle, so named because carbon cycles from CO2 to glucose. Perhaps a better name for this cycle would be the “energy cycle”. The point of photosynthesis is to use the sun’s energy to make ATP, which is then used to make glucose (which stores the energy). The point of cellular respiration is to use the energy from glucose to make ATP, which is used to drive work in the cell. Photosynthetic organisms, such as plants, algae, and some bacteria, can do both parts of the carbon cycle. Non-photosynthetic organisms can only do half. The next two chapters will describe the two halves of the carbon cycle in detail.

10.4.2 Redox Reactions and Energy
Free energy is defined as the ability to do work. Energy exists in different forms, including electrical energy, light energy, chemical energy, and heat energy. In order to appreciate the way energy flows into and out of biological systems, it is important to understand more about the different types of energy that exist in the physical world.
Energy production within a cell involves many coordinated biochemical pathways. Most of these pathways are combinations of oxidation and reduction reactions. Oxidation and reduction occur in tandem. An oxidation reaction strips an electron from an atom in a compound, and then adds this electron to another compound in a reduction reaction. Because oxidation and reduction usually occur together, these pairs of reactions are called reduction-oxidation, or redox reactions. A mnemonic to help remember which is which is OIL RIG, which stands for Oxidation Is Loss of an electron and Reduction Is Gain of an electron.
The removal of an electron from a molecule, oxidizing it, results in a decrease in potential energy in the oxidized compound. The transfer of electrons between molecules is important because most of the chemical energy stored in molecules is in the form of high-energy electrons. The transfer of energy in the form of electrons allows the cell to transfer and use energy in an incremental fashion—in small packages rather than in a single, destructive burst.
10.4.3 Electron Carriers
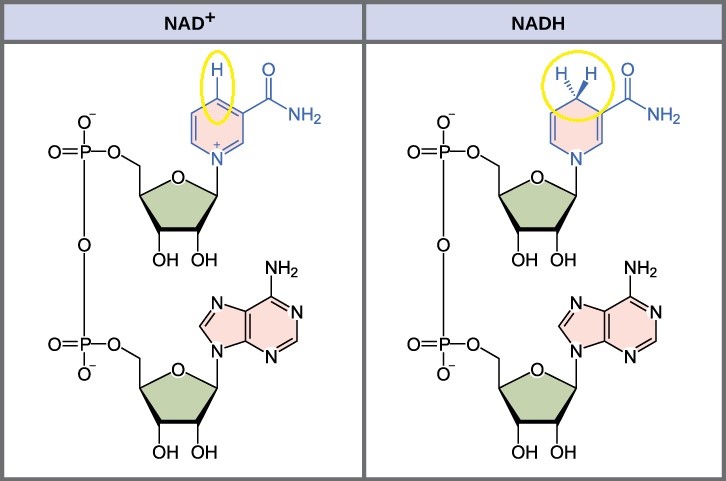
In living systems, a small class of compounds functions as electron shuttles: They bind and carry high-energy electrons between compounds in pathways. The principal electron carriers we will consider are derived from the B vitamin group and are derivatives of nucleotides. These compounds can be easily reduced (by accepting electrons) or oxidized (by losing electrons). Nicotinamide Adenine Dinucleotide (NAD+) (Figure 10.16) is derived from vitamin B3 (niacin). NAD+ is the oxidized form of the molecule; NADH is the reduced form of the molecule after it has accepted two electrons and a proton (the equivalent of a hydrogen atom plus an extra electron).
NAD+ can accept electrons from an organic molecule according to the general equation:
|
R-H |
+ |
NAD+ |
|
NADH |
+ |
R |
|
Reducing Agent |
|
Oxidizing Agent |
|
Reduced Form |
|
Oxidized Form |
Similarly, flavin adenine dinucleotide (FAD) is derived from vitamin B2 (riboflavin). Its reduced form is FADH2. NADP+ is a variant of NAD+ that has an extra phosphate group. Both NAD+ and FAD are used in energy extraction from sugars, and NADP+ plays an important role in photosynthesis.
10.5 Metabolic Pathways
Learning Objectives
By the end of this section, you will be able to:
- Define a metabolic pathway and identify whether one is catabolic or anabolic.
- Discuss feedback inhibition in metabolic pathways.
A metabolic pathway is a series of sequential biochemical reactions that convert a substrate molecule or molecules, step-by-step, through a series of metabolic intermediates, eventually yielding a final product or products. Pathways that build more complex molecules out of smaller ones are called anabolic pathways. Pathways that break more complex molecules into smaller ones are called catabolic pathways. Anabolic pathways require energy, while catabolic pathways produce energy. Consequently, metabolism is composed of building (anabolism) and degradation (catabolism) (Figure 10.17).
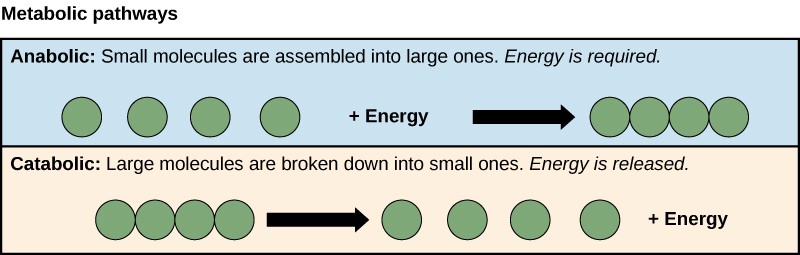
10.4.1 Evolution of Metabolic Pathways
There is more to the complexity of metabolism than understanding the metabolic pathways alone. Metabolic complexity varies from organism to organism. Photosynthesis is the primary pathway in which photosynthetic organisms like plants harvest the sun’s energy and convert it into carbohydrates. (The majority of global synthesis is done by planktonic algae.) The by-product of photosynthesis is oxygen, required by some cells to carry out cellular respiration. During cellular respiration, oxygen aids in the catabolic breakdown of carbon compounds, like carbohydrates. Among the products of this catabolism are CO2 and ATP. In addition, some eukaryotes perform catabolic processes without oxygen (fermentation); that is, they perform anaerobic metabolism.
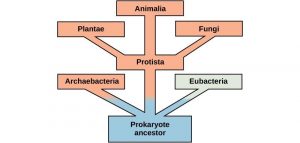
Organisms probably evolved anaerobic metabolism to survive, since living organisms came into existence about 3.8 billion years ago, when the atmosphere lacked oxygen. Despite the differences between organisms and the complexity of metabolism, researchers have found that all branches of life share some of the same metabolic pathways, suggesting that all organisms evolved from the same ancient common ancestor (Figure 10.18). Evidence indicates that over time, the pathways diverged, adding specialized enzymes to allow organisms to better adapt to their environment, thus increasing their chance to survive. However, the underlying principle remains that all organisms must harvest energy from their environment and convert it to ATP to carry out cellular functions.
10.4.2 Feedback Inhibition in Metabolic Pathways
Molecules can regulate enzyme function in many ways. Perhaps the most relevant sources of enzyme regulatory molecules are the products of the cellular metabolic reactions themselves. In a most efficient and elegant way, cells have evolved to use the products of their own reactions for feedback inhibition of enzyme activity. Feedback inhibition involves the use of a reaction product to regulate its own further production (Figure 10.19). The cell responds to the abundance of specific products by slowing down production. Such reaction products may inhibit the enzymes that catalyzed their production through the mechanisms described above.
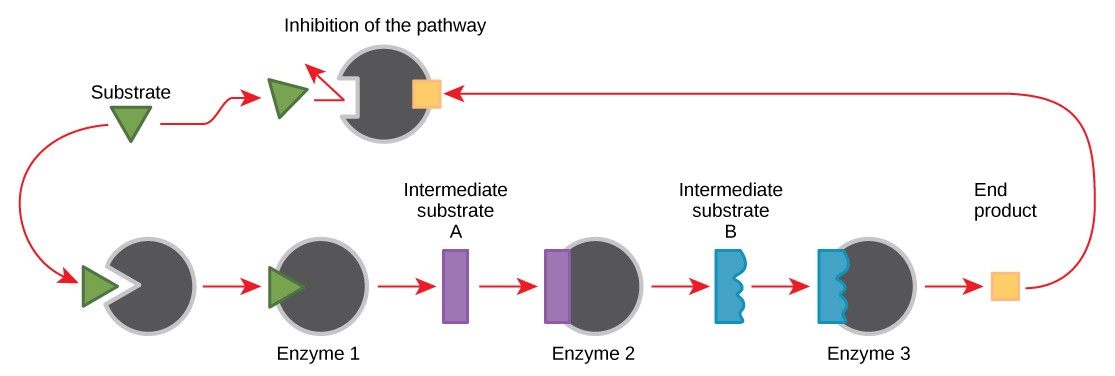
For example, ATP is an allosteric regulator of some of the enzymes involved in the catabolic breakdown of sugar, the process that produces ATP. In this way, when ATP is abundant, the cell can prevent its further production. Remember that ATP is an unstable molecule that can spontaneously dissociate into ADP. If too much ATP were present in a cell, much of it would go to waste. On the other hand, ADP serves as an allosteric activator for some of the same enzymes that are inhibited by ATP. Thus, when relative levels of ADP are high compared to ATP, the cell is triggered to produce more ATP.

